The Gaussian discriminate analysis is a generative learning algorithm which models the PDF using a (multivariate) Gaussian distribution. This class of density estimation is also called parametric density estimation.
Parametric Density Estimation: To get an idea of the underlying distribution of the features we try to estimate this distribution based on a formula with parameters. Most commenly used formula for parametric density estimation is the Gaussian distribution:
1D Gaussian
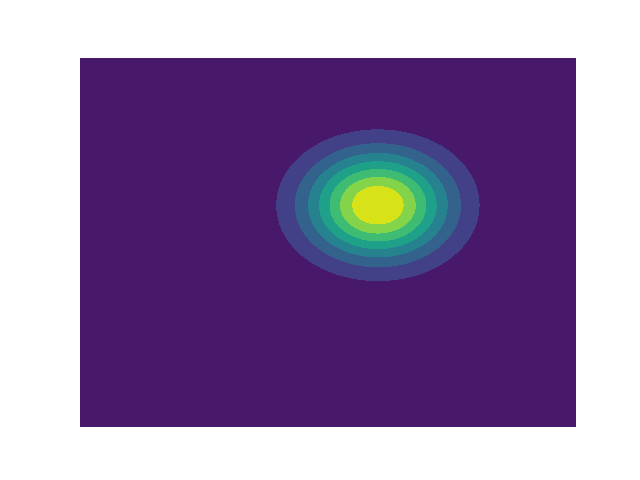
Multivariate Gaussian
Quadratic discriminate analysis
Quadratic discriminate analysis is a form of GDA where the class conditional probability of each class is modeled using individual gaussians, which means that $\sigma$ of each class are not necessarily the same.
In binary (two class) QDA the classification is based on the discriminant:
A positive discriminant means the posterior probability of class 1 is higher and thus we classify $x$ as class 1. The log of the posterior probabilities is taken because this yields a simpler (more efficient) equation. This is possible because log is a monotonically increasing function, which means that the relations of order are unchanged. The $max_i$ $p(y_i|x)$ will also be the $max_i$ log $p(y_i|x)$. From the log of the posterior probability follows:
Removing all non-class dependent constants yields:
Plugging the result into the discriminant equation yields:
This is a quadratic equation modeling the decision boundary for a two class classification problem. The final class prediction or hypothesis $h(x)$ is then given by the following bernouilli distribution.
Linear discriminate analysis
Given the assumption that the covariance matrices of both classes are equal $(\Sigma_1 == \Sigma_2)$, the decision boundary becomes linear:
Also called Fisher's Linear Discriminate analysis.
LDA example
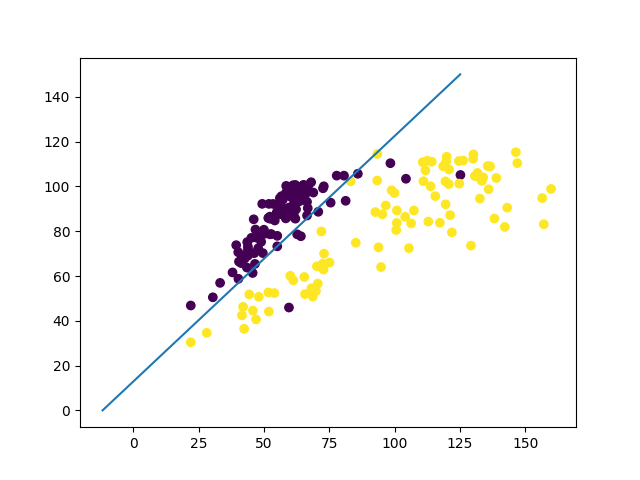
LDA as a dimensionality reduction technique
In Fisher's article he observes that at the decision boundary (discriminant = 0), the ratio of between and within class variance is:
Here $\omega$ is a unit vector in a certain direction. He furthermore obeserves that this ratio is maximal for, $\omega$ as in LDA:
The simple interpretation of this is that, the dicision boundary has a slope which is equal to the direction of maximum $\Phi$.
The use LDA as a dimensionality reduction technique all you have to do is map the data onto the first $k$ principal axes of the $\Phi$. (Later on we will have a look at other similar dimensionality reduction technique such as PCA)